Recently I attended an inspirational tech talk hosted by Databricks where Denny Lee shared some great tips and techniques around analyzing COVID-19 Open Research Dataset (CORD-19) freely available here. As stated in its description:
“In response to the COVID-19 pandemic, the White House and a coalition of leading research groups have prepared the COVID-19 Open Research Dataset (CORD-19). CORD-19 is a resource of over 44,000 scholarly articles, including over 29,000 with full text, about COVID-19, SARS-CoV-2, and related coronaviruses.”
In this blog post, I’m sharing some of the analysis presented during the tech talk which I have now replicated using StreamSets Platform. In particular, I’ve created the following the data pipeline tool in StreamSets Transformer running on Databricks cluster.
While this blog addresses a sensitive subject, the goal is to show how to use some of the tools you might have at your disposal in order to better understand and analyze the problem at hand.
Ok, let’s get started.
JSON to Parquet
The first pipeline reads a subset of CORD-19 data available in JSON format, repartitions the data (where the number of partitions is set to number of nodes in the cluster), and converts the data into Parquet for efficient downstream processing in Apache Spark. (Note: the data in my case just happens to be stored on Amazon S3, but in your case it may be stored on other sources as well.)
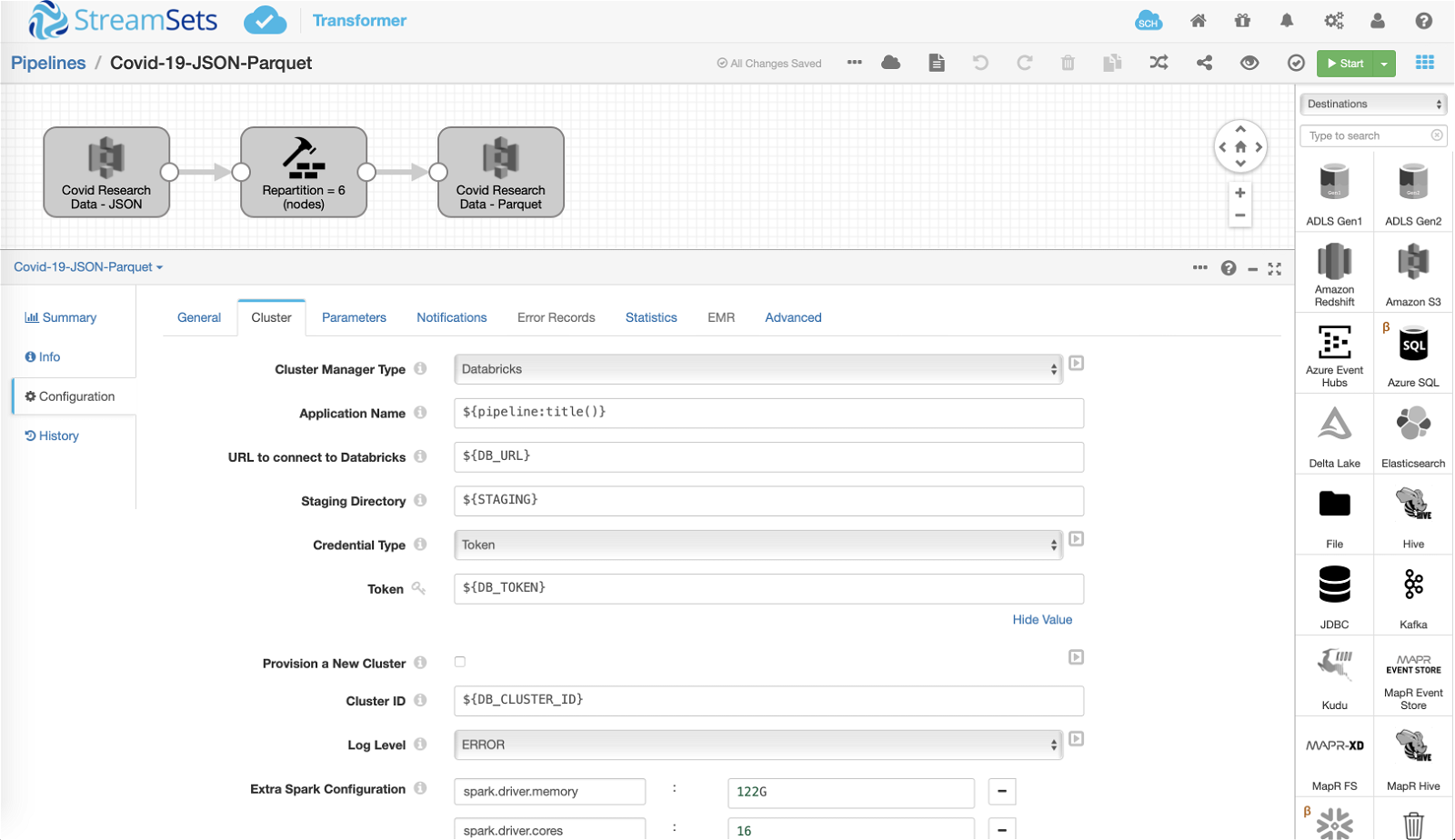
Delta Lake
The second pipeline reads the CORD-19 data stored in Parquet format, performs some transformations, and stores the curated data on Databricks File System (DBFS) for further analysis.
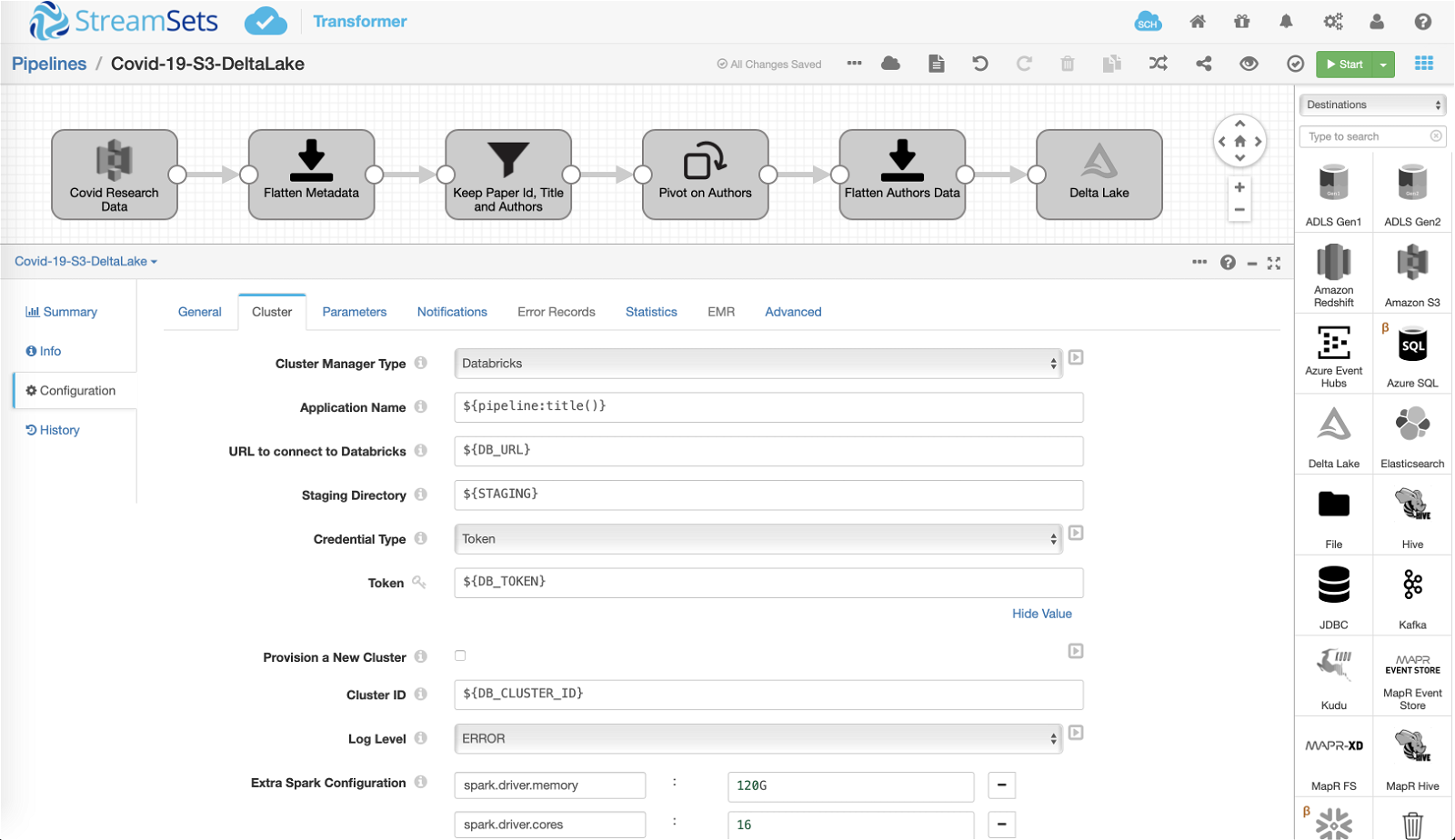
One of the key transformations that occur in this pipeline is “pivot” – the CORD-19 dataset is structured in a way where each research paper (record) contains a nested list of affiliated authors. So in order to analyze individual author records, we need to create one output record for each author in the nested list.
Databricks Notebook
Once the transformed data was available on DBFS, I created a Databricks Notebook to visualize geo locations of the authors — basically showing the total number of research papers produced by authors from each country.
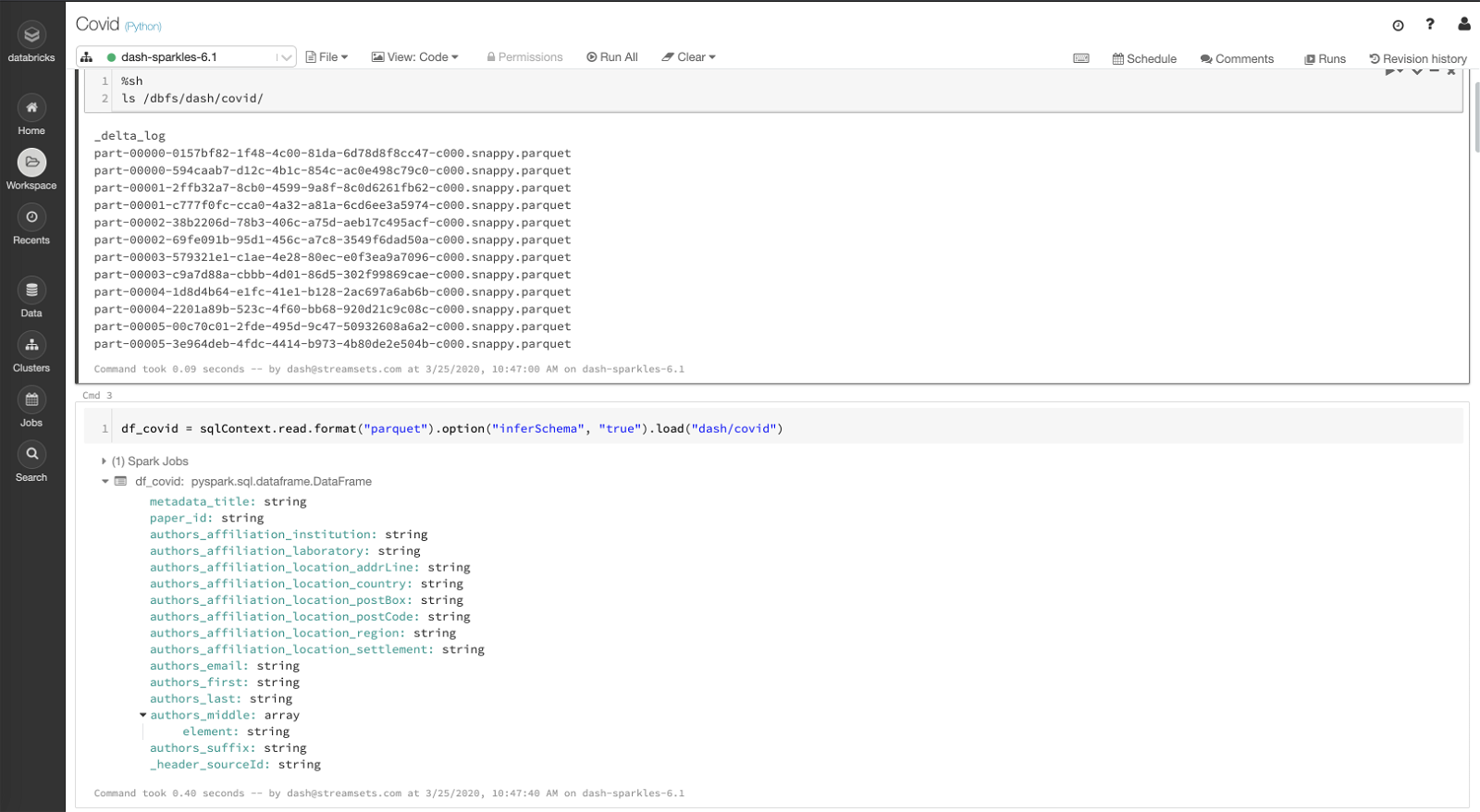
Here I am showing the transformed data on DBFS — list of Parquet files and the dataframe schema that was generated/created when the data was written out to DBFS by Delta Lake destination in the second pipeline above.
Here the first query creates a temp table paper_by_country which holds paper_id and the author’s country affiliated with it based on authors_affiliation_location_country column. The second query uses the temp table and joins it with country_mapping to count the number of papers and plots the location on the map.
paper_by_country = spark.sql("""select paper_id,min(authors_affiliation_location_country) as author_country from covid
where authors_affiliation_location_country is NOT NULL
group by paper_id""")
paper_by_country.createOrReplaceTempView("paper_by_country")
select m.Alpha3, count(distinct p.paper_id) as number_of_papers from paper_by_country p left join country_mapping m on m.AuthorCountry = p.author_country group by m.Alpha3
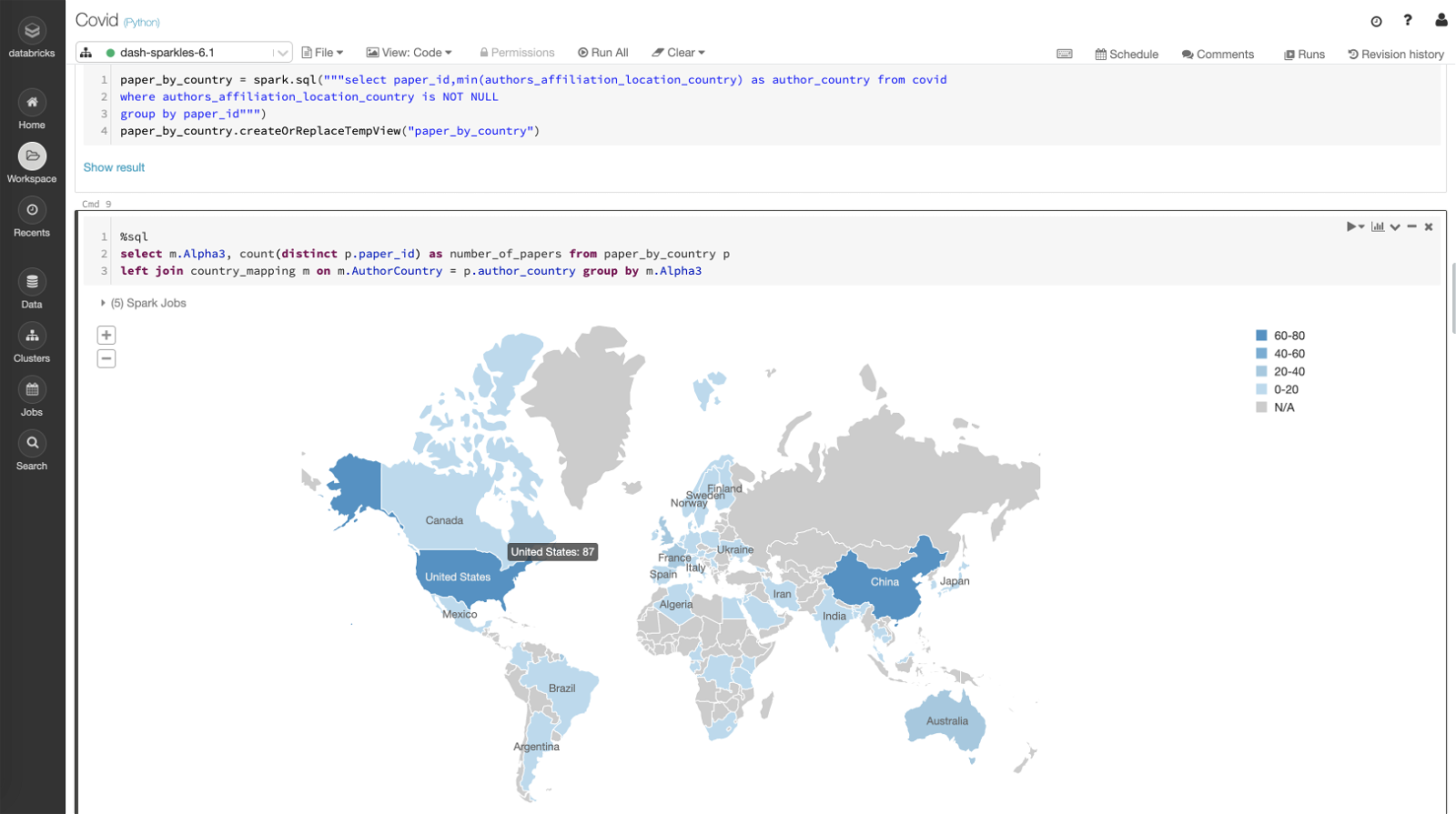
Notes:
- I’ve analyzed a small subset of the CORD-19 dataset so the numbers shown in any queries, results, map, etc. are not a true representation of the entire dataset.
- The country code mapping (“country_mapping“) I’ve used in the above SQL has been put together by one of the Databricks presenters and should be available on their webinar page. (If I find it, I will provide a link to it here.)
Conclusion
As you can imagine, I haven’t even scratched the surface and there’s ample opportunity for exploring and analyzing the CORD-19 dataset. You can also participate in this Kaggle competition to win prizes and, more importantly, help the community by generating new insights. Hopefully this blog post gives you some ideas, tips and tricks to get started.
Learn more about StreamSets for Databricks which is available on Microsoft Azure Marketplace and AWS Marketplace.

